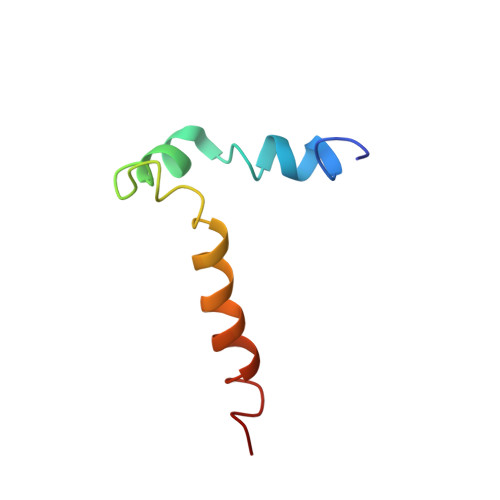
Structural and Functional Analysis of the Transmembrane Segment Pair VI and VII of the NHE1 Isoform of the Na(+)/H(+) Exchanger.
Alves, C., Lee, B.L., Sykes, B.D., Fliegel, L.(2014) Biochemistry 53: 3658-3670
- PubMed: 24840010
- DOI: https://doi.org/10.1021/bi500392y
- Primary Citation of Related Structures:
2MDF - PubMed Abstract:
Isoform 1 of the mammalian Na(+)/H(+) exchanger (NHE1) is a ubiquitously expressed plasma membrane pH regulatory protein. It removes one intracellular H(+) in exchange for one extracellular Na(+). The 500 N-terminal amino acids comprise the catalytic membrane domain and fold into 12 transmembrane (TM) segments. To gain insight into the structure and function of human NHE1, a region spanning transmembrane domains VI and VII was expressed and purified, and the structure was determined using nuclear magnetic resonance (NMR). Segment VI includes two structurally conserved regions corresponding to two short α-helices involving residues 229-236 and 239-247. Segment VII includes one long helical region spanning residues 255-274. The NMR structure of the peptide containing transmembrane domains VI and VII was very similar to the previously published structures of the single-transmembrane segments except that TM VII was not kinked. Tryptophan scanning site-directed mutagenesis of TM VI demonstrated that mutation of residues V240-V245 to tryptophan eliminated NHE1 activity when the full length protein was expressed in cells. In contrast, mutants F246W and E247W were functional. Double mutant V242F/F260V retained activity, while the individual mutations were not active. The results suggest that the region of TM VI from V240 to V245 is closely associated with TM VII and that, in agreement with the NMR structure of VI-VII segments, V242 and F260 are in close association. A study of two transmembrane peptides provides further insight into the structure of the NHE1 protein.
Organizational Affiliation:
Department of Biochemistry, University of Alberta , Edmonton, Alberta, Canada T6G 2H7.