Abstract
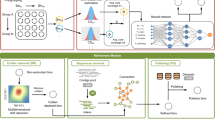
Assembling metagenomic data sequenced by NGS platforms poses significant computational challenges, especially due to large volumes of data, sequencing errors, and variations in size, complexity, diversity and abundance of organisms present in a given metagenome. To overcome these problems, this work proposes an open-source, bioinformatic tool called GCSplit, which partitions metagenomic sequences into subsets using a computationally inexpensive metric: the GC content. Experiments performed on real data show that preprocessing short reads with GCSplit prior to assembly reduces memory consumption and generates higher quality results, such as an increase in the size of the largest contig and N50 metric, while both the L50 value and the total number of contigs produced in the assembly were reduced. GCSplit is available at https://github.com/mirand863/gcsplit.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Vogel, T.M., Simonet, P., Jansson, J.K., et al.: TerraGenome: a consortium for the sequencing of a soil metagenome. Nat. Rev. Microbiol. 7, 252 (2009)
Venter, J.C., Remington, K., Heidelberg, J.F., et al.: Environmental genome shotgun sequencing of the Sargasso Sea. Science 304, 66–74 (2004)
Qin, J., Li, R., Raes, J., et al.: A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010)
Turnbaugh, P.J., Ley, R.E., Hamady, M., et al.: The human microbiome project: exploring the microbial part of ourselves in a changing world. Nature 449, 804–810 (2007)
Namiki, T., Hachiya, T., Tanaka, H., et al.: MetaVelvet: an extension of Velvet assembler to De Novo metagenome assembly from short sequence reads. Nucleic Acids Res. 40, e155 (2012)
Rodrigue, S., Materna, A.C., Timberlake, S., et al.: Unlocking short read sequencing for metagenomics. PLoS ONE 5, e11840 (2010)
Nielsen, H.B., Almeida, M., Juncker, A.S., et al.: Identification and assembly of genomes and genetic elements in complex metagenomic samples without using reference genomes. Nat. Biotechnol. 32, 822–828 (2014)
Wojcieszek, M., Pawełkowicz, M., Nowak, R., et al.: Genomes correction and assembling: present methods and tools. In: SPIE Proceedings, vol. 9290, p. 92901X (2014)
Charuvaka, A., Rangwala, H.: Evaluation of short read metagenomic assembly. BMC Genom. 12, S8 (2011)
Rasheed, Z., Rangwala, H.: Mc-MinH: metagenome clustering using minwise based hashing. In: SIAM International Conference in Data Mining, pp. 677–685 (2013)
Howe, A.C., Jansson, J.K., Malfatti, S.A., et al.: Tackling soil diversity with the assembly of large, complex metagenomes. Proc. Natl. Acad. Sci. 111, 4904–4909 (2014)
Nurk, S., Meleshko, D., Korobeynikov, A., et al.: metaSPAdes: a new versatile metagenomic assembler. Genome Res. 27, 824–834 (2017)
Brown, C.T., Howe, A., Zhang, Q., et al.: A reference-free algorithm for computational normalization of shotgun sequencing data. arXiv:1203.4802 (2012)
Haas, B.J., Papanicolaou, A., Yassour, M., et al.: De Novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 8, 1494–1512 (2013)
McCorrison, J.M., Venepally, P., Singh, I., et al.: NeatFreq: reference-free data reduction and coverage normalization for De Novo sequence assembly. BMC bioinform. 15, 357 (2014)
Durai, D.A., Schulz, M.H.: In-silico read normalization using set multi-cover optimization. bioRxiv:133579 (2017)
Pell, J., Hintze, A., Canino-Koning, R., et al.: Scaling metagenome sequence assembly with probabilistic de Bruijn graphs. Proc. Natl. Acad. Sci. 109, 13272–13277 (2012)
Crusoe, M.R., Alameldin, H.F., Awad, S., et al.: The khmer software package: enabling efficient nucleotide sequence analysis. F1000Research 4, 900 (2015)
Rengasamy, V., Medvedev, P., Madduri, K.: Parallel and memory-efficient preprocessing for metagenome assembly. In: IPDPSW, pp. 283–292 (2017)
Cleary, B., Brito, I.L., Huang, K., et al.: Detection of low-abundance bacterial strains in metagenomic datasets by eigengenome partitioning. Nat. Biotechnol. 33, 1053–1060 (2015)
Melsted, P., Halldórsson, B.V.: KmerStream: streaming algorithms for k-mer abundance estimation. Bioinformatics 30, 3541–3547 (2014)
Bankevich, A., Nurk, S., Antipov, D., et al.: SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012)
Stamps, B.W., Corsetti, F.A., Spear, J.R., et al.: Draft genome of a novel Chlorobi member assembled by tetranucleotide binning of a hot spring metagenome. Genome Announc. 2, e00897–e00914 (2014)
Ibarbalz, F.M., Orellana, E., Figuerola, E.L., et al.: Shotgun metagenomic profiles have a high capacity to discriminate samples of activated sludge according to wastewater type. Appl. Environ. Microbiol. 82, 5186–5196 (2016)
Gurevich, A., Saveliev, V., Vyahhi, N., et al.: QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013)
Acknowledgments
This research is supported in part by CNPq under grant numbers 421528/2016–8 and 304711/2015–2. The authors would also like to thank CAPES for granting scholarships. Datasets processed in Sagarana HPC cluster, CPAD–ICB–UFMG.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG, part of Springer Nature
About this paper
Cite this paper
Miranda, F., Batista, C., Silva, A., Morais, J., Neto, N., Ramos, R. (2018). Improving Metagenomic Assemblies Through Data Partitioning: A GC Content Approach. In: Rojas, I., Ortuño, F. (eds) Bioinformatics and Biomedical Engineering. IWBBIO 2018. Lecture Notes in Computer Science(), vol 10813. Springer, Cham. https://doi.org/10.1007/978-3-319-78723-7_36
Download citation
DOI: https://doi.org/10.1007/978-3-319-78723-7_36
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-78722-0
Online ISBN: 978-3-319-78723-7
eBook Packages: Computer ScienceComputer Science (R0)