Quorum sensing
This article needs additional citations for verification. (April 2024) |
In biology, quorum sensing or quorum signaling (QS)[1] is the process of cell-to-cell communication[2] that allows bacteria to detect and respond to cell population density by gene regulation, typically as a means of acclimating to environmental disadvantages.[3]
More specifically, quorum sensing is a type of cellular signaling, and more specifically can be considered a type of paracrine signaling. However, it also contains traits of autocrine signaling: a cell produces both an autoinducer molecule and the receptor for the autoinducer.[3] As one example, QS enables bacteria to restrict the expression of specific genes to the high cell densities at which the resulting phenotypes will be most beneficial, especially for phenotypes that would be ineffective at low cell densities and therefore too energetically costly to express.[4] Many species of bacteria use quorum sensing to coordinate gene expression according to the density of their local population. In a similar fashion, some social insects use quorum sensing to determine where to nest. Quorum sensing in pathogenic bacteria activates host immune signaling and prolongs host survival, by limiting the bacterial intake of nutrients, such as tryptophan, which further is converted to serotonin.[5] As such, quorum sensing allows a commensal interaction between host and pathogenic bacteria.[5] Quorum sensing may also be useful for cancer cell communications.[6]
In addition to its function in biological systems, quorum sensing has several useful applications for computing and robotics. In general, quorum sensing can function as a decision-making process in any decentralized system in which the components have: (a) a means of assessing the number of other components they interact with and (b) a standard response once a threshold to achieve number of components is useful for amino acid regulation is detected.
Discovery
[edit]The first observations of an autoinducer-controlled phenotype in bacteria were reported in 1970, by Kenneth Nealson, Terry Platt, and J. Woodland Hastings,[7] who observed what they described as a conditioning of the medium in which they had grown the bioluminescent marine bacterium Aliivibrio fischeri.[8] These bacteria did not synthesize luciferase—and therefore did not luminesce—in freshly inoculated culture but only after the bacterial population had increased significantly.
Etymology
[edit]Because Nealson, Platt, and Hastings attributed the conditioning of the growth medium to the growing population of cells itself, they referred to the phenomenon as autoinduction.[7][9][8] In 1994, after study of the phenomenon had expanded into several additional bacteria, Stephen Winans did not believe the word autoinduction fully characterized the true process so, in a review article coauthored with W. Claiborne Fuqua and E. Peter Greenberg,[10] he introduced the term quorum sensing. Its use also avoided confusion between the terms autoinduction and autoregulation.
The new term was not stumbled onto, but rather created through trial and error. Among the alternatives that Winans had created and considered were gridlockins, communiolins, and quoromones.[11]
Bacteria
[edit]| Part of a series on |
| Microbial and microbot movement |
|---|
 |
| Microswimmers |
| Molecular motors |

Some of the best-known examples of quorum sensing come from studies of bacteria. Bacteria use quorum sensing to regulate certain phenotype expressions, which in turn, coordinate their behaviors. Some common phenotypes include biofilm formation, virulence factor expression, and motility. Certain bacteria are able to use quorum sensing to regulate bioluminescence, nitrogen fixation and sporulation.[12]
The quorum-sensing function is based on the local density of the bacterial population in the immediate environment.[13] It can occur within a single bacterial species, as well as between diverse species. Both gram-positive and gram-negative bacteria use quorum sensing, but there are some major differences in their mechanisms.[14]
Mechanism
[edit]For the bacteria to use quorum sensing constitutively, they must possess three abilities: secretion of a signaling molecule, secretion of an autoinducer (to detect the change in concentration of signaling molecules), and regulation of gene transcription as a response.[12] This process is highly dependent on the diffusion mechanism of the signaling molecules. QS signaling molecules are usually secreted at a low level by individual bacteria. At low cell density, the molecules may just diffuse away. At high cell density, the local concentration of signaling molecules may exceed its threshold level, and trigger changes in gene expression.[14]
Gram-positive bacteria
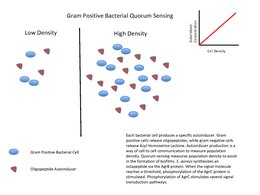
[edit]Gram-positive bacteria use autoinducing peptides (AIP) as their autoinducers.[15]
When gram-positive bacteria detect high concentration of AIPs in their environment, that happens by way of AIPs binding to a receptor to activate a kinase. The kinase phosphorylates a transcription factor, which regulates gene transcription. This is called a two-component system.
Another possible mechanism is that AIP is transported into the cytosol, and binds directly to a transcription factor to initiate or inhibit transcription.[15]
Gram-negative bacteria
[edit]Gram-negative bacteria produce N-acyl homoserine lactones (AHL) as their signaling molecule.[15] Usually AHLs do not need additional processing, and bind directly to transcription factors to regulate gene expression.[14]
Some gram-negative bacteria may use the two-component system as well.[15]
Examples
[edit]Aliivibrio fischeri
[edit]The bioluminescent bacterium Aliivibrio fischeri is the first organism in which QS was observed. It lives as a mutualistic symbiont in the photophore (or light-producing organ) of the Hawaiian bobtail squid. When A. fischeri cells are free-living (or planktonic), the autoinducer is at low concentration, and, thus, cells do not show luminescence. However, when the population reaches the threshold in the photophore (about 1011 cells/ml), transcription of luciferase is induced, leading to bioluminescence. In A. fischeri, bioluminescence is regulated by AHLs (N-acyl-homoserine lactones) which is a product of the LuxI gene whose transcription is regulated by the LuxR activator. LuxR works only when AHLs binds to the LuxR.
Curvibacter sp.
[edit]Curvibacter sp. is a gram-negative curved rod-formed bacterium which is the main colonizer of the epithelial cell surfaces of the early branching metazoan Hydra vulgaris.[16][17] Sequencing the complete genome uncovered a circular chromosome (4.37 Mb), a plasmid (16.5 kb), and two operons coding each for an AHL (N-acyl-homoserine lactone) synthase (curI1 and curI2) and an AHL receptor (curR1 and curR2).[17] Moreover, a study showed that these host associated Curvibacter bacteria produce a broad spectrum of AHL, explaining the presence of those operons.[17] As mentioned before, AHL are the quorum sensing molecules of gram-negative bacteria, which means Curvibacter has a quorum sensing activity.
Even though their function in host-microbe interaction is largely unknown, Curvibacter quorum-sensing signals are relevant for host-microbe interactions.[17] Indeed, due to the oxidoreductase activity of Hydra, there is a modification of AHL signalling molecules (3-oxo-homoserine lactone into 3-hydroxy-homoserine lactone) which leads to a different host-microbe interaction. On one hand, a phenotypic switch of the colonizer Curvibacter takes place. The most likely explanation is that the binding of 3-oxo-HSL and 3-hydroxy-HSL causes different conformational changes in the AHL receptors curR1 and curR2. As a result, there is a different DNA-binding motif affinity and thereby different target genes are activated.[17] On the other hand, this switch modifies its ability to colonize the epithelial cell surfaces of Hydra vulgaris.[17] Indeed, one explanation is that with a 3-oxo-HSL quorum-sensing signal, there is an up-regulation of flagellar assembly. Yet, flagellin, the main protein component of flagella, can act as an immunomodulator and activate the innate immune response in Hydra. Therefore, bacteria have less chance to evade the immune system and to colonize host tissues.[17] Another explanation is that 3-hydroxy-HSL induces carbon metabolism and fatty acid degradation genes in Hydra. This allows the bacterial metabolism to adjust itself to the host growth conditions, which is essential for the colonization of the ectodermal mucus layer of Hydrae.[17]
Enterococcus faecalis
[edit]Enterococcus faecalis is an opportunistic, gram-positive bacteria that forms biofilm in glass. This process is also known as forming a biofilm in vitro. The presence of (Esp), a certain cell surface protein, aids the formation of a biofilm by E. faecalis.[18]
The ability of E. faecalis to form biofilms contributes to its capacity to survive in extreme environments, and facilitates its involvement in persistent bacterial infection, particularly in the case of multi-drug resistant strains.[19] Biofilm formation in E. faecalis is associated with DNA release, and such release has emerged as a fundamental aspect of biofilm formation.[19] Conjugative plasmid DNA transfer in E. faecalis is enhanced by the release of peptide sex pheromones.[20]
Escherichia coli
[edit]In the gram-negative bacterium Escherichia coli, cell division may be partially regulated by AI-2-mediated quorum sensing. This species uses AI-2, which is produced and processed by the lsr operon. Part of it encodes an ABC transporter, which imports AI-2 into the cells during the early stationary (latent) phase of growth. AI-2 is then phosphorylated by the LsrK kinase, and the newly produced phospho-AI-2 can be either internalized or used to suppress LsrR, a repressor of the lsr operon (thereby activating the operon). Transcription of the lsr operon is also thought to be inhibited by dihydroxyacetone phosphate (DHAP) through its competitive binding to LsrR. Glyceraldehyde 3-phosphate has also been shown to inhibit the lsr operon through cAMP-CAPK-mediated inhibition. This explains why, when grown with glucose, E. coli will lose the ability to internalize AI-2 (because of catabolite repression). When grown normally, AI-2 presence is transient.
E. coli and Salmonella enterica do not produce AHL signals commonly found in other gram-negative bacteria. However, they have a receptor that detects AHLs from other bacteria and change their gene expression in accordance with the presence of other "quorate" populations of gram-negative bacteria.[21] AHL quorum sensing regulates a wide range of genes through cell density. Other species of bacteria produce AHLs that Escherichia and Salmonella can detect. E. coli and Salmonella produce a receptor like protein (SdiA) allowing the amino acid sequence that is similar to AHL show AHLs can be found in the bovine rumen and E. coli responds to AHLs taken out of the bovine rumen. Most animals do not have AHL in their gastrointestinal tracts.[22]
Salmonella enterica
[edit]Salmonella encodes a LuxR homolog, SdiA, but does not encode an AHL synthase. SdiA detects AHLs produced by other species of bacteria including Aeromonas hydrophila, Hafnia alvei, and Yersinia enterocolitica.[23] When AHL is detected, SdiA regulates the rck operon on the Salmonella virulence plasmid (pefI-srgD-srgA-srgB-rck-srgC) and a single gene horizontal acquisition in the chromosome srgE.[24][25] Salmonella does not detect AHL when passing through the gastrointestinal tracts of several animal species, suggesting that the normal microbiota does not produce AHLs. However, SdiA does become activated when Salmonella transits through turtles colonized with Aeromonas hydrophila or mice infected with Yersinia enterocolitica.[26][27] Therefore, Salmonella appears to use SdiA to detect the AHL production of other pathogens rather than the normal gut flora.
Myxococcus xanthus
[edit]Myxococcus is a genus of gram-negative bacterium in the Myxococcacae family. Myxococcus xanthus specifically, a bacillus myxobacteria species within Myxococcae, grows in the upper layers of soil. This bacterium is known for its unique utilization of quorum sensing practices to hunt.
The bacterium uniquely survives not on sugars, but lipids created by the degradation of macromolecules lysed by the species. It hunts and feeds through a density-regulated method of predation that is "the regulation of gene expression in response to cell density."[28] The pilus propelled microorganism moves with the use of both S- and A- (or gliding) motility, which provide transportation across a dynamic range of different surfaces.[29] M. xanthus's A-motility is most effective in the presence of a single or low number of cells, allowing the bacteria to glide in high agar concentrations. The S-motility, or social motility, is controlled by the process of quorum sensing and is only effective when cells are within one cell length of a neighbor.[30] Although the precise specifics of M. xanthus's communication methods for quorum sensing are not well understood, the bacteria mediate the process by using both C-signal and A-factor. The A-factor molecule, produced by M. xanthus, must reach a set concentration to initiate aggregation for hunting.[31] The C-signal concentration, on the other hand, plays a role in fruiting body production.
The species is known for its ability to use quorum sensing to hunt in special packs with thousands of individual cells, lending to M. xanthus's name "the wolf packs." M. xanthus is inclined to behave in a multicellular fashion. In the presence of many cells, it uses these "wolf packs" to form "highly structured biofilms that include tentacle-like packs of surface-gliding cell groups, synchronized rippling waves of oscillating cells and massive spore-filled aggregates that protrude upwards from the substratum to form fruiting bodies."[32][28] On the fringes of this film, individual cells can be observed "gliding across the surface, but the majority of cells are observed in large tendril-shaped groups" using S-motility.[28]
Staphylococcus aureus
[edit]Staphylococcus aureus is a type of pathogen that causes infection to the skin and soft tissue and can lead to a variety of more severe diseases such as osteomyelitis, pneumonia, and endocarditis. S. aureus uses biofilms in order to increase its chances of survival by becoming resistant to antibiotics. Biofilms help S. aureus become up to 1500 times more resistant to antibiofilm agents, which try to break down biofilms formed by S. aureus.[33]
Pseudomonas aeruginosa
[edit]The environmental bacterium and opportunistic pathogen Pseudomonas aeruginosa uses quorum sensing to coordinate the formation of biofilm, swarming motility, exopolysaccharide production, virulence, and cell aggregation.[34] These bacteria can grow within a host without harming it until they reach a threshold concentration. Then they become aggressive, developing to the point at which their numbers are sufficient to overcome the host's immune system, and form a biofilm, leading to disease within the host as the biofilm is a protective layer encasing the bacterial population.[citation needed] The relative ease of growth, handling, and genetic manipulation of P. aeruginosa has lent much research effort to the quorum sensing circuits of this relatively common bacterium. Quorum sensing in P. aeruginosa typically encompasses two complete AHL synthase-receptor circuits, LasI-LasR and RhlI-RhlR, as well as the orphan receptor-regulator QscR, which is also activated by the LasI-generated signal.[35] Together, the multiple AHL quorum sensing circuits of P. aeruginosa influence regulation of hundreds of genes.
Another form of gene regulation that allows the bacteria to rapidly adapt to surrounding changes is through environmental signaling. Recent studies have discovered that anaerobiosis can significantly impact the major regulatory circuit of quorum sensing. This important link between quorum sensing and anaerobiosis has a significant impact on the production of virulence factors of this organism.[36] There is hope among some humans that the therapeutic enzymatic degradation of the signaling molecules will be possible when treating illness caused by biofilms, and prevent the formation of such biofilms and possibly weaken established biofilms. Disrupting the signaling process in this way is called quorum sensing inhibition.[37]
Acinetobacter sp.
[edit]It has recently been found that Acinetobacter sp. also show quorum sensing activity. This bacterium, an emerging pathogen, produces AHLs.[38] Acinetobacter sp. shows both quorum sensing and quorum quenching activity. It produces AHLs and can also degrade the AHL molecules.[38]
Aeromonas sp.
[edit]This bacterium was previously considered a fish pathogen, but it has recently emerged as a human pathogen.[39] Aeromonas sp. have been isolated from various infected sites from patients (bile, blood, peritoneal fluid, pus, stool and urine). All isolates produced the two principal AHLs, N-butanoylhomoserine lactone (C4-HSL) and N-hexanoyl homoserine lactone (C6-HSL). It has been documented that Aeromonas sobria has produced C6-HSL and two additional AHLs with N-acyl side chain longer than C6.[40]
Yersinia
[edit]The YenR and YenI proteins produced by the gammaproteobacterium Yersinia enterocolitica are similar to Aliivibrio fischeri LuxR and LuxI.[41][42] YenR activates the expression of a small non-coding RNA, YenS. YenS inhibits YenI expression and acylhomoserine lactone production.[43] YenR/YenI/YenS are involved in the control of swimming and swarming motility.[42][43]
Molecules involved
[edit]Three-dimensional structures of proteins involved in quorum sensing were first published in 2001, when the crystal structures of three LuxS orthologs were determined by X-ray crystallography.[44] In 2002, the crystal structure of the receptor LuxP of Vibrio harveyi with its inducer AI-2 (which is one of the few biomolecules containing boron) bound to it was also determined.[45] Many bacterial species, including E. coli, an enteric bacterium and model organism for gram-negative bacteria, produce AI-2. A comparative genomic and phylogenetic analysis of 138 genomes of bacteria, archaea, and eukaryotes found that "the LuxS enzyme required for AI-2 synthesis is widespread in bacteria, while the periplasmic binding protein LuxP is present only in Vibrio strains," leading to the conclusion that either "other organisms may use components different from the AI-2 signal transduction system of Vibrio strains to sense the signal of AI-2 or they do not have such a quorum sensing system at all."[46] Vibrio species utilize Qrr RNAs, small non-coding RNAs, that are activated by these autoinducers to target cell density master regulators. Farnesol is used by the fungus Candida albicans as a quorum sensing molecule that inhibits filamentation.[47]
A database of quorum-sensing peptides is available under the name Quorumpeps.[48][49]
Certain bacteria can produce enzymes called lactonases that can target and inactivate AHLs. Researchers have developed novel molecules which block the signalling receptors of bacteria ("Quorum quenching"). mBTL is a compound that has been shown to inhibit quorum sensing and decrease the amount of cell death by a significant amount.[50] Additionally, researchers are also examining the role of natural compounds (such as caffeine) as potential quorum sensing inhibitors.[51] Research in this area has been promising and could lead to the development of natural compounds as effective therapeutics.
Evolution
[edit]Sequence analysis
[edit]The majority of quorum sensing systems that fall under the "two-gene" (an autoinducer synthase coupled with a receptor molecule) paradigm as defined by the Vibrio fischeri system occur in the gram-negative Pseudomonadota. A comparison between the Pseudomonadota phylogeny as generated by 16S ribosomal RNA sequences and phylogenies of LuxI-, LuxR-, or LuxS-homologs shows a notably high level of global similarity. Overall, the quorum sensing genes seem to have diverged along with the Pseudomonadota phylum as a whole. This indicates that these quorum sensing systems are quite ancient, and arose very early in the Pseudomonadota lineage.[52][53]
LuxI and LuxR have coevolved through a long history of horizontal gene transfer (HGT) events. An early study reconciling their gene trees with the rRNA tree suggested frequent HGT events for both LuxI and LuxR, indicating that they are horizontally transferred together and coevolve due to their functional dependency.[52] Similarly, in QS systems in bacteria associated with Populus deltoides, the gene trees for luxI and luxR show high topological similarity, indicating coevolution of cognate pairs.[54] In addition to horizontal transfer of complete LuxI/LuxR-type QS systems, many Proteobacteria genomes exhibit an excess of LuxR genes or cases with only LuxR but not LuxI, acquired from different sources via HGT.[54] Due to the frequent transfer of functional pairs of homologs (i.e., LuxI/LuxR-type systems from multiple independent sources), it is possible that the regulatory hierarchy formed by the LuxI/LuxR and RhlR-RhlI systems is a result of sequential integration of circuits obtained from different sources, due to interactions between multiple homologs.[52] Interestingly, LuxI genes have likely undergone horizontal gene transfer from Proteobacteria to other lineages, as they have been detected in Nitrospira lineage II.[55]
In quorum sensing genes of Gammaproteobacteria, which includes Pseudomonas aeruginosa and Escherichia coli, the LuxI/LuxR genes form a functional pair, with LuxI as the auto-inducer synthase and LuxR as the receptor. Gammaproteobacteria are unique in possessing quorum sensing genes, which, although functionally similar to the LuxI/LuxR genes, have a markedly divergent sequence.[53] This family of quorum-sensing homologs may have arisen in the Gammaproteobacteria ancestor, although the cause of their extreme sequence divergence yet maintenance of functional similarity has yet to be explained. In addition, species that employ multiple discrete quorum sensing systems are almost all members of the Gammaproteobacteria, and evidence of horizontal transfer of quorum sensing genes is most evident in this class.[52][53]
Interaction of quorum-sensing molecules with mammalian cells and its medical applications
[edit]Next to the potential antimicrobial functionality, quorum-sensing derived molecules, especially the peptides, are being investigated for their use in other therapeutic domains as well, including immunology, central nervous system disorders and oncology. Quorum-sensing peptides have been demonstrated to interact with cancer cells, as well as to permeate the blood–brain barrier reaching the brain parenchyma.[56][57][58]
Role of quorum sensing in biofilm development
[edit]Quorum sensing (QS) is used by bacteria to form biofilms. Quorum sensing is used by bacteria to form biofilms because the process determines if the minimum number of bacteria necessary for biofilm formation are present. The criteria to form a biofilm is dependent on a certain density of bacteria rather than a certain number of bacteria being present. When aggregated in high enough densities, some bacteria may form biofilms to protect themselves from biotic or abiotic threats.[59] Quorum sensing is used by both Gram-positive and Gram-negative bacteria because it aids cellular reproduction. Once in a biofilm, bacteria can communicate with other bacteria of the same species. Bacteria can also communicate with other species of bacteria. This communication is enabled through autoinducers used by the bacteria.[13]
Additionally, certain responses can be generated by the host organism in response to the certain bacterial autoinducers. Despite the fact that specific bacterial quorum sensing systems are different, for example the target genes, signal relay mechanisms, and chemical signals used between bacteria, the ability to coordinate gene expression for a specific species of bacteria remains the same. This ability alludes to the larger idea that bacteria have potential to become a multicellular bacterial body.[13]
Secondly, biofilms may also serve to transport nutrients into the microbial community or transport toxins out by means of channels that permeate the extracellular polymeric matrix (like cellulose) that holds the cells together. Finally, biofilms are an ideal environment for horizontal gene transfer through either conjugation or environmental DNA (eDNA) that exists in the biofilm matrix.[59]
The process of biofilm development is often triggered by environmental signals, and bacteria are proven to require flagella to successfully approach a surface, adhere to it, and form the biofilm.[59] As cells either replicate or aggregate in a location, the concentration of autoinducers outside of the cells increases until a critical mass threshold is reached. At this point, it is energetically unfavorable for intracellular autoinducers to leave the cell and they bind to receptors and trigger a signaling cascade to initiate gene expression and begin secreting an extracellular polysaccharide to encase themselves inside.[60]
One modern method of preventing biofilm development without the use of antibiotics is with anti-QS substances, such (naringenin, taxifolin, etc) that can be utilized as alternative form of therapy against bacterial virulence.[61]
Archaea
[edit]Examples
[edit]Methanosaeta harundinacea 6Ac
[edit]Methanosaeta harundinacea 6Ac, a methanogenic archaeon, produces carboxylated acyl homoserine lactone compounds that facilitate the transition from growth as short cells to growth as filaments.[62]
Viruses
[edit]A mechanism involving arbitrium has recently been described in bacteriophages infecting several Bacillus species.[63][64] The viruses communicate with each other to ascertain their own density compared to potential hosts. They use this information to decide whether to enter a lytic or lysogenic life-cycle.[65] This decision is crucial as it affects their replication strategy and potential to spread within the host population, optimizing their survival and proliferation under varying environmental conditions. This communication mechanism enables a coordinated infection strategy, significantly enhancing the efficiency of phage proliferation. By synchronizing their life cycles, bacteriophages can maximize their impact on the host population, potentially leading to more effective control of bacterial densities.
Plants
[edit]QS is important to plant-pathogen interactions, and their study has also contributed to the QS field more generally.[66][8] The first X-ray crystallography results for some of the key proteins were those of Pantoea stewartii subsp. stewartii in maize/corn[67][8] and Agrobacterium tumefaciens, a crop pathogen with a wider range of hosts.[68][69][8] These interactions are facilitated by quorum-sensing molecules and play a major role in maintaining the pathogenicity of bacteria towards other hosts, such as humans. This mechanism can be understood by looking at the effects of N-Acyl homoserine lactone (AHL), one of the quorum sensing-signaling molecules in gram-negative bacteria, on plants. The model organism used is Arabidopsis thaliana.[70] Further insights reveal that AHLs influence plant immune responses and can alter plant hormone levels, thereby affecting plant growth and susceptibility to infection. Understanding these dynamics is crucial for developing innovative strategies to combat plant diseases and improve agricultural productivity. Researchers have also noted that certain plants can degrade these signaling molecules, potentially as a defensive strategy to disrupt bacterial communication. This interplay between bacterial signaling and plant responses suggests a complex co-evolutionary relationship that could be exploited to enhance crop resistance to bacterial pathogens.
The role of AHLs having long carbon-chains (C12, C14), which have an unknown receptor mechanism, is less well understood than AHLs having short carbon-chains (C4, C6, C8), which are perceived by the G protein-coupled receptor. A phenomenon called "AHL priming", which is a dependent signalling pathway, enhanced our knowledge of long-chain AHLs. The role of quorum-sensing molecules was better explained according to three categories: host physiology–based impact of quorum sensing molecules; ecological effects; and cellular signaling. Calcium signalling and calmodulin have a large role in short-chain AHLs' response in Arabidopsis. Research was also conducted on barley and the crop called yam bean (Pachyrhizus erosus) that reveals the AHLs determining the detoxification enzymes called GST were found less in yam bean.[71]
Quorum sensing-based regulatory systems are necessary to plant-disease-causing bacteria. Looking towards developing new strategies based on plant-associated microbiomes, the aim of further study is to improve the quantity and quality of the food supply. Further research into this inter-kingdom communication also enhances the possibility of learning about quorum sensing in humans.[72] This exploration could open new avenues for managing microbial communities in agricultural settings, potentially leading to the development of more sustainable farming practices that leverage natural microbial processes to boost crop resilience and productivity.
Quorum quenching
[edit]Quorum quenching is the process of preventing quorum sensing by disrupting signalling.[73] This is achieved by inactivating signalling enzymes, by introducing molecules that mimic signalling molecules and block their receptors, by degrading signalling molecules themselves, or by a modification of the quorum sensing signals due to an enzyme activity.[17][73][74][75]
Inhibition
[edit]Closantel and triclosan are known inhibitors of quorum sensing enzymes.[76] Closantel induces aggregation of the histidine kinase sensor in two-component signalling. The latter disrupts the synthesis of a class of signalling molecules known as N-acyl homoserine lactones (AHLs) by blocking the enoyl-acyl carrier protein (ACP) reductase.[76][77]
Mimicry
[edit]Two groups of well-known mimicking molecules include halogenated furanones, which mimic AHL molecules, and synthetic Al peptides (AIPs), which mimic naturally occurring AIPs. These groups inhibit receptors from binding substrate or decrease the concentration of receptors in the cell.[76] Furanones have also been found to act on AHL-dependant transcriptional activity, whereby the half life of the autoinducer-binding LuxR protein is significantly shortened.[78]
Degradation
[edit]Recently, a well-studied quorum quenching bacterial strain (KM1S) was isolated and its AHL degradation kinetics were studied using rapid resolution liquid chromatography (RRLC).[79] RRLC efficiently separates components of a mixture to a high degree of sensitivity, based on their affinities for different liquid phases.[80] It was found that the genome of this strain encoded an inactivation enzyme with distinct motifs targeting the degradation of AHLs.[79]
Modifications
[edit]As mentioned before, N-acyl-homoserine lactones (AHL) are the quorum sensing signaling molecules of the gram-negative bacteria. However, these molecules may have different functional groups on their acyl chain, and also a different length of acyl chain. Therefore, there exist many different AHL signaling molecules, for example, 3-oxododecanoyl-L-homoserine lactone (3OC12-HSL) or 3-hydroxydodecanoyl-L-homoserine lactone (3OHC12-HSL). The modification of those quorum sensing (QS) signaling molecules is another sort of quorum quenching. This can be carried out by an oxidoreductase activity.[17] As an example, we will discuss the interaction between a host, Hydra vulgaris, and the main colonizer of its epithelial cell surfaces, Curvibacter spp. Those bacteria produce 3-oxo-HSL quorum sensing molecules.[17] However, the oxidoreductase activity of the polyp Hydra is able to modify the 3-oxo-HSL into their 3-hydroxy-HSL counterparts.[17] We can characterize this as quorum quenching since there is an interference with quorum sensing molecules. In this case, the outcomes differ from simple QS inactivation: the host modification results in a phenotypic switch of Curvibacter, which modifies its ability to colonize the epithelial cell surfaces of H. vulgaris.[17]
Applications
[edit]Applications of quorum quenching that have been exploited by humans include the use of AHL-degrading bacteria in aquacultures to limit the spread of diseases in aquatic populations of fish, mollusks and crustaceans.[81] This technique has also been translated to agriculture, to restrict the spread of pathogenic bacteria that use quorum sensing in plants.[81][82] Anti-biofouling is another process that exploits quorum quenching bacteria to mediate the dissociation of unwanted biofilms aggregating on wet surfaces, such as medical devices, transportation infrastructure and water systems.[81][83] Quorum quenching is recently studied for the control of fouling and emerging contaminants in electro membrane bioreactors (eMBRs) for the advanced treatment of wastewater.[84] Extracts of several traditional medicinal herbs display quorum quenching activity, and have potential antibacterial applications.[85][86]
Social insects
[edit]Social insect colonies are an excellent example of a decentralized system, because no individual is in charge of directing or making decisions for the colony. Several groups of social insects have been shown to use quorum sensing in a process that resembles collective decision-making.
Examples
[edit]Ants
[edit]Colonies of the ant Temnothorax albipennis nest in small crevices between rocks. When the rocks shift and the nest is broken up, these ants must quickly choose a new nest to move into. During the first phase of the decision-making process, a small portion of the workers leave the destroyed nest and search for new crevices. When one of these scout ants finds a potential nest, she assesses the quality of the crevice based on a variety of factors including the size of the interior, the number of openings (based on light level), and the presence or absence of dead ants.[87][88] The worker then returns to the destroyed nest, where she waits for a short period before recruiting other workers to follow her to the nest that she has found, using a process called tandem running. The waiting period is inversely related to the quality of the site; for instance, a worker that has found a poor site will wait longer than a worker that encountered a good site.[89] As the new recruits visit the potential nest site and make their own assessment of its quality, the number of ants visiting the crevice increases. During this stage, ants may be visiting many different potential nests. However, because of the differences in the waiting period, the number of ants in the best nest will tend to increase at the greatest rate. Eventually, the ants in this nest will sense that the rate at which they encounter other ants has exceeded a particular threshold, indicating that the quorum number has been reached.[90] Once the ants sense a quorum, they return to the destroyed nest and begin rapidly carrying the brood, queen, and fellow workers to the new nest. Scouts that are still tandem-running to other potential sites are also recruited to the new nest, and the entire colony moves. Thus, although no single worker may have visited and compared all of the available options, quorum sensing enables the colony as a whole to quickly make good decisions about where to move.
Honey bees
[edit]Honey bees (Apis mellifera) also use quorum sensing to make decisions about new nest sites. Large colonies reproduce through a process called swarming, in which the queen leaves the hive with a portion of the workers to form a new nest elsewhere. After leaving the nest, the workers form a swarm that hangs from a branch or overhanging structure. This swarm persists during the decision-making phase until a new nest site is chosen.
The quorum sensing process in honey bees is similar to the method used by Temnothorax ants in several ways. A small portion of the workers leave the swarm to search out new nest sites, and each worker assesses the quality of the cavity it finds. The worker then returns to the swarm and recruits other workers to her cavity using the honey bee waggle dance. However, instead of using a time delay, the number of dance repetitions the worker performs is dependent on the quality of the site. Workers that found poor nests stop dancing sooner, and can, therefore, be recruited to the better sites. Once the visitors to a new site sense that a quorum number (usually 10–20 bees) has been reached, they return to the swarm and begin using a new recruitment method called piping. This vibration signal causes the swarm to take off and fly to the new nest location. In an experimental test, this decision-making process enabled honey bee swarms to choose the best nest site in four out of five trials.[91][92]
Synthetic biology
[edit]Quorum sensing has been engineered using synthetic biological circuits in different systems. Examples include rewiring the AHL components to toxic genes to control population size in bacteria;[93] and constructing an auxin-based system to control population density in mammalian cells.[94] Synthetic quorum sensing circuits have been proposed to enable applications like controlling biofilms[95] or enabling drug delivery.[96] Quorum sensing based genetic circuits have been used to convert AI-2 signals to AI-1 and then subsequently use the AI-1 signal to alter bacterial growth rate, thereby changing the composition of a consortium.[97]
Remarkable advancements have been and are continuing to be made in recent years in our understanding of synthetic biology in terms of endocrine and paracrine signaling mechanisms, and the myriad of modes by which bacteria record domestic and foreign cell numbers.[98] The modulation of gene expression in response to oscillations in cell-population density is thanks to the QS techniques regulating bacterial communication natural and artificial cultures. It is also clear that intra- and inter-species cell–cell communication occurs and is regulated by quorum sensing systems. Further, there is mounting data demonstrating that autoinducer signals elicit specific responses from eukaryotic hosts.
Computing and robotics
[edit]Quorum sensing can be a useful tool for improving the function of self-organizing networks such as the SECOAS (Self-Organizing Collegiate Sensor) environmental monitoring system. In this system, individual nodes sense that there is a population of other nodes with similar data to report. The population then nominates just one node to report the data, resulting in power savings.[99] Ad hoc wireless networks can also benefit from quorum sensing, by allowing the system to detect and respond to network conditions.[100]
Quorum sensing can also be used to coordinate the behavior of autonomous robot swarms. Using a process similar to that used by Temnothorax ants, robots can make rapid group decisions without the direction of a controller.[101]
Despite recent advancements, the true nature of these back-and-forth conversations remains a mystery, and further rigorous research targeting inter- and intra- species communication is still necessary to maximize knowledge of quorum sensing and its potential to improve research and treatments of cancer and bacterial diseases. The code to understanding these complex bacterial languages is to decipher the impact of the words.[98]
See also
[edit]References
[edit]- ^ Lupp C, Urbanowski M, Greenberg EP, Ruby EG (October 2003). "The Vibrio fischeri quorum-sensing systems ain and lux sequentially induce luminescence gene expression and are important for persistence in the squid host". Molecular Microbiology. 50 (1). Wiley Publishing: 319–331. doi:10.1046/j.1365-2958.2003.t01-1-03585.x. PMID 14507383.
- ^ Rutherford, S. T.; Bassler, B. L. (2012). "Bacterial Quorum Sensing: Its Role in Virulence and Possibilities for Its Control". Cold Spring Harbor Perspectives in Medicine. 2 (11): a012427. doi:10.1101/cshperspect.a012427. PMC 3543102. PMID 23125205.
- ^ a b Postat, J; Bousso, P (September 11, 2019). "Quorum Sensing by Monocyte-Derived Populations". Frontiers in Immunology. 10 (2140): 2140. doi:10.3389/fimmu.2019.02140. PMC 6749007. PMID 31572366.
- ^ Papenfort, K; Bassler, B (February 11, 2017). "Quorum-Sensing Signal-Response Systems in Gram-Negative Bacteria". Nature Reviews Microbiology. 14 (9): 576–588. doi:10.1038/nrmicro.2016.89. PMC 5056591. PMID 27510864.
- ^ a b Jugder BE, Batista JH, Gibson JA, Cunningham PM, Asara JM, Watnick PI (September 2022). "Vibrio cholerae high cell density quorum sensing activates the host intestinal innate immune response". Cell Reports. 40 (12): 111368. doi:10.1016/j.celrep.2022.111368. PMC 9534793. PMID 36130487.
- ^ Ali I, Alfarouk KO, Reshkin SJ, Ibrahim ME (16 January 2018). "Doxycycline as Potential Anti-cancer Agent". Anti-Cancer Agents in Medicinal Chemistry. 17 (12): 1617–1623. doi:10.2174/1871520617666170213111951. PMID 28270076.
- ^ a b Nealson KH, Platt T, Hastings JW (October 1970). "Cellular control of the synthesis and activity of the bacterial luminescent system". Journal of Bacteriology. 104 (1): 313–322. doi:10.1128/jb.104.1.313-322.1970. PMC 248216. PMID 5473898.
- ^ a b c d e Von Bodman SB, Bauer WD, Coplin DL (2003). "Quorum sensing in plant-pathogenic bacteria". Annual Review of Phytopathology. 41 (1). Annual Reviews: 455–482. doi:10.1146/annurev.phyto.41.052002.095652. PMID 12730390. S2CID 3926735.
- ^ Nealson KH (February 1977). "Autoinduction of bacterial luciferase. Occurrence, mechanism and significance". Archives of Microbiology. 112 (1). Springer Science and Business Media LLC: 73–79. Bibcode:1977ArMic.112...73N. doi:10.1007/bf00446657. PMID 843170. S2CID 34239721.
- ^ Fuqua, W C; Winans, S C; Greenberg, E P (January 1994). "Quorum sensing in bacteria: the LuxR-LuxI family of cell density-responsive transcriptional regulators". Journal of Bacteriology. 176 (2): 269–275. doi:10.1128/jb.176.2.269-275.1994. PMC 205046. PMID 8288518.
- ^ Turovskiy, Yevgeniy; Kashtanov, Dimitri; Paskhover, Boris; Chikindas, Michael L. (2007). "Quorum Sensing: Fact, Fiction, and Everything in Between". Advances in Applied Microbiology Volume 62. Vol. 62. pp. 191–234. doi:10.1016/S0065-2164(07)62007-3. ISBN 978-0-12-373669-7. PMC 2391307. PMID 17869606.
{{cite book}}:|journal=ignored (help) - ^ a b Pan J, Ren D (November 2009). "Quorum sensing inhibitors: a patent overview". Expert Opinion on Therapeutic Patents. 19 (11): 1581–1601. doi:10.1517/13543770903222293. PMID 19732032. S2CID 30007165.
- ^ a b c Miller, Melissa B.; Bassler, Bonnie L. (October 2001). "Quorum Sensing in Bacteria". Annual Review of Microbiology. 55 (1): 165–199. doi:10.1146/annurev.micro.55.1.165. PMID 11544353.
- ^ a b c Bassler BL (December 1999). "How bacteria talk to each other: regulation of gene expression by quorum sensing". Current Opinion in Microbiology. 2 (6): 582–587. doi:10.1016/s1369-5274(99)00025-9. PMID 10607620.
- ^ a b c d Rutherford ST, Bassler BL (November 2012). "Bacterial quorum sensing: its role in virulence and possibilities for its control". Cold Spring Harbor Perspectives in Medicine. 2 (11): a012427. doi:10.1101/cshperspect.a012427. PMC 3543102. PMID 23125205.
- ^ "Curvibacter fontana sp. nov., a microaerobic bacteria isolated from well water". ResearchGate. Retrieved 2019-03-13.
- ^ a b c d e f g h i j k l m Pietschke C, Treitz C, Forêt S, Schultze A, Künzel S, Tholey A, et al. (October 2017). "Host modification of a bacterial quorum-sensing signal induces a phenotypic switch in bacterial symbionts". Proceedings of the National Academy of Sciences of the United States of America. 114 (40): E8488–E8497. Bibcode:2017PNAS..114E8488P. doi:10.1073/pnas.1706879114. PMC 5635886. PMID 28923926.
- ^ Kristich, C. J.; Li, Y. H.; Cvitkovitch, D. G.; Dunny, G. M. (2004). "Esp-Independent Biofilm Formation by Enterococcus faecalis". Journal of Bacteriology. 186 (1): 154–163. doi:10.1128/JB.186.1.154-163.2004. PMC 365672. PMID 14679235.
- ^ a b Șchiopu P, Toc DA, Colosi IA, Costache C, Ruospo G, Berar G, Gălbău ȘG, Ghilea AC, Botan A, Pană AG, Neculicioiu VS, Todea DA (July 2023). "An Overview of the Factors Involved in Biofilm Production by the Enterococcus Genus". Int J Mol Sci. 24 (14): 11577. doi:10.3390/ijms241411577. PMC 10380289. PMID 37511337.
- ^ Hirt H, Greenwood-Quaintance KE, Karau MJ, Till LM, Kashyap PC, Patel R, Dunny GM (February 2018). "Enterococcus faecalis Sex Pheromone cCF10 Enhances Conjugative Plasmid Transfer In Vivo". mBio. 9 (1). doi:10.1128/mBio.00037-18. PMC 5821081. PMID 29440568.
- ^ Ahmer BM (May 2004). "Cell-to-cell signalling in Escherichia coli and Salmonella enterica". Molecular Microbiology. 52 (4): 933–945. doi:10.1111/j.1365-2958.2004.04054.x. PMID 15130116.
- ^ Soares, J. A.; Ahmer, B. M. (2011). "Detection of acyl-homoserine lactones by Escherichia and Salmonella". Current Opinion in Microbiology. 14 (2): 188–193. doi:10.1016/j.mib.2011.01.006. PMC 3078957. PMID 21353625.
- ^ Michael B, Smith JN, Swift S, Heffron F, Ahmer BM (October 2001). "SdiA of Salmonella enterica is a LuxR homolog that detects mixed microbial communities". Journal of Bacteriology. 183 (19): 5733–5742. doi:10.1128/JB.183.19.5733-5742.2001. PMC 95466. PMID 11544237.
- ^ Ahmer BM, van Reeuwijk J, Timmers CD, Valentine PJ, Heffron F (March 1998). "Salmonella typhimurium encodes an SdiA homolog, a putative quorum sensor of the LuxR family, that regulates genes on the virulence plasmid". Journal of Bacteriology. 180 (5): 1185–1193. doi:10.1128/JB.180.5.1185-1193.1998. PMC 107006. PMID 9495757.
- ^ Smith JN, Ahmer BM (February 2003). "Detection of other microbial species by Salmonella: expression of the SdiA regulon". Journal of Bacteriology. 185 (4): 1357–1366. doi:10.1128/JB.185.4.1357-1366.2003. PMC 142872. PMID 12562806.
- ^ Smith JN, Dyszel JL, Soares JA, Ellermeier CD, Altier C, Lawhon SD, et al. (July 2008). Ausubel, Frederick M. (ed.). "SdiA, an N-acylhomoserine lactone receptor, becomes active during the transit of Salmonella enterica through the gastrointestinal tract of turtles". PLOS ONE. 3 (7): e2826. Bibcode:2008PLoSO...3.2826S. doi:10.1371/journal.pone.0002826. PMC 2475663. PMID 18665275.
- ^ Dyszel JL, Smith JN, Lucas DE, Soares JA, Swearingen MC, Vross MA, et al. (January 2010). "Salmonella enterica serovar Typhimurium can detect acyl homoserine lactone production by Yersinia enterocolitica in mice". Journal of Bacteriology. 192 (1): 29–37. doi:10.1128/JB.01139-09. PMC 2798265. PMID 19820103.
- ^ a b c Berleman, James E.; Kirby, John R. (2009-09-01). "Deciphering the hunting strategy of a bacterial wolfpack". FEMS Microbiology Reviews. 33 (5): 942–957. doi:10.1111/j.1574-6976.2009.00185.x. ISSN 0168-6445. PMC 2774760. PMID 19519767. Retrieved 2024-04-16.
- ^ Shi, W; Zusman, D R (1993-04-15). "The two motility systems of Myxococcus xanthus show different selective advantages on various surfaces". Proceedings of the National Academy of Sciences of the United States of America. 90 (8): 3378–3382. Bibcode:1993PNAS...90.3378S. doi:10.1073/pnas.90.8.3378. ISSN 0027-8424. PMC 46303. PMID 8475084.
- ^ Patra, Pintu; Kissoon, Kimberley; Cornejo, Isabel; Kaplan, Heidi B.; Igoshin, Oleg A. (2016-06-30). "Colony Expansion of Socially Motile Myxococcus xanthus Cells Is Driven by Growth, Motility, and Exopolysaccharide Production". PLOS Computational Biology. 12 (6): –1005010. Bibcode:2016PLSCB..12E5010P. doi:10.1371/journal.pcbi.1005010. ISSN 1553-7358. PMC 4928896. PMID 27362260.
- ^ Lloyd, Daniel G.; Whitworth, David E. (2017-03-14). "The Myxobacterium Myxococcus xanthus Can Sense and Respond to the Quorum Signals Secreted by Potential Prey Organisms". Frontiers in Microbiology. 8: 439. doi:10.3389/fmicb.2017.00439. ISSN 1664-302X. PMC 5348527. PMID 28352265.
- ^ Dye, Keane J.; Yang, Zhaomin (2022-04-29). "Analysis of Myxococcus xanthus Vegetative Biofilms With Microtiter Plates". Frontiers in Microbiology. 13: 894562. doi:10.3389/fmicb.2022.894562. ISSN 1664-302X. PMC 9100584. PMID 35572678.
- ^ Wu, Xiying; Wang, Huan; Xiong, Juan; Yang, Guo-Xun; Hu, Jin-Feng; Zhu, Quangang; Chen, Zhongjian (June 2024). "Staphylococcus aureus biofilm: Formulation, regulatory, and emerging natural products-derived therapeutics". Biofilm. 7: 100175. doi:10.1016/j.bioflm.2023.100175. PMC 10827693. PMID 38298832.
- ^ Sauer K, Camper AK, Ehrlich GD, Costerton JW, Davies DG (February 2002). "Pseudomonas aeruginosa displays multiple phenotypes during development as a biofilm". Journal of Bacteriology. 184 (4): 1140–1154. doi:10.1128/jb.184.4.1140-1154.2002. PMC 134825. PMID 11807075.
- ^ Schuster, Martin; Peter Greenberg, E. (April 2006). "A network of networks: Quorum-sensing gene regulation in Pseudomonas aeruginosa". International Journal of Medical Microbiology. 296 (2–3): 73–81. doi:10.1016/j.ijmm.2006.01.036. PMID 16476569.
- ^ Cornelis, P., ed. (2008). Pseudomonas: Genomics and Molecular Biology (1st ed.). Caister Academic Press. ISBN 978-1-904455-19-6. Retrieved 1 May 2022.
- ^ Ma, Yeping; Tang, Wing Suet; Liu, Sylvia Yang; Khoo, Bee Luan; Chua, Song Lin (9 February 2024). "Juglone as a Natural Quorum Sensing Inhibitor against Pseudomonas aeruginosa pqs-Mediated Virulence and Biofilms". ACS Pharmacology & Translational Science. 7 (2): 533–543. doi:10.1021/acsptsci.3c00354. PMC 10863437. PMID 38357290.
- ^ a b Chan KG, Atkinson S, Mathee K, Sam CK, Chhabra SR, Cámara M, et al. (March 2011). "Characterization of N-acylhomoserine lactone-degrading bacteria associated with the Zingiber officinale (ginger) rhizosphere: co-existence of quorum quenching and quorum sensing in Acinetobacter and Burkholderia". BMC Microbiology. 11 (1): 51. doi:10.1186/1471-2180-11-51. PMC 3062576. PMID 21385437.
- ^ Igbinosa IH, Igumbor EU, Aghdasi F, Tom M, Okoh AI (2012). "Emerging Aeromonas species infections and their significance in public health". TheScientificWorldJournal. 2012: 625023. doi:10.1100/2012/625023. PMC 3373137. PMID 22701365.
- ^ Chan KG, Puthucheary SD, Chan XY, Yin WF, Wong CS, Too WS, Chua KH (January 2011). "Quorum sensing in Aeromonas species isolated from patients in Malaysia". Current Microbiology. 62 (1): 167–172. doi:10.1007/s00284-010-9689-z. PMID 20544198. S2CID 6761810.
- ^ Throup JP, Camara M, Briggs GS, Winson MK, Chhabra SR, Bycroft BW, et al. (July 1995). "Characterisation of the yenI/yenR locus from Yersinia enterocolitica mediating the synthesis of two N-acylhomoserine lactone signal molecules". Molecular Microbiology. 17 (2): 345–356. doi:10.1111/j.1365-2958.1995.mmi_17020345.x. PMID 7494483. S2CID 3100775.
- ^ a b Atkinson S, Chang CY, Sockett RE, Cámara M, Williams P (February 2006). "Quorum sensing in Yersinia enterocolitica controls swimming and swarming motility". Journal of Bacteriology. 188 (4): 1451–1461. doi:10.1128/JB.188.4.1451-1461.2006. PMC 1367215. PMID 16452428.
- ^ a b Tsai CS, Winans SC (April 2011). "The quorum-hindered transcription factor YenR of Yersinia enterocolitica inhibits pheromone production and promotes motility via a small non-coding RNA". Molecular Microbiology. 80 (2): 556–571. doi:10.1111/j.1365-2958.2011.07595.x. PMID 21362062.
- ^ Lewis HA, Furlong EB, Laubert B, Eroshkina GA, Batiyenko Y, Adams JM, et al. (June 2001). "A structural genomics approach to the study of quorum sensing: crystal structures of three LuxS orthologs". Structure. 9 (6): 527–537. doi:10.1016/S0969-2126(01)00613-X. PMID 11435117.
- ^ Chen X, Schauder S, Potier N, Van Dorsselaer A, Pelczer I, Bassler BL, Hughson FM (January 2002). "Structural identification of a bacterial quorum-sensing signal containing boron". Nature. 415 (6871): 545–549. doi:10.1038/415545a. PMID 11823863.
- ^ Sun J, Daniel R, Wagner-Döbler I, Zeng AP (September 2004). "Is autoinducer-2 a universal signal for interspecies communication: a comparative genomic and phylogenetic analysis of the synthesis and signal transduction pathways". BMC Evolutionary Biology. 4 (1): 36. doi:10.1186/1471-2148-4-36. PMC 524169. PMID 15456522.
- ^ Hornby JM, Jensen EC, Lisec AD, Tasto JJ, Jahnke B, Shoemaker R, et al. (July 2001). "Quorum sensing in the dimorphic fungus Candida albicans is mediated by farnesol". Applied and Environmental Microbiology. 67 (7): 2982–2992. Bibcode:2001ApEnM..67.2982H. doi:10.1128/AEM.67.7.2982-2992.2001. PMC 92970. PMID 11425711.
- ^ Wynendaele E, Bronselaer A, Nielandt J, D'Hondt M, Stalmans S, Bracke N, et al. (January 2013). "Quorumpeps database: chemical space, microbial origin and functionality of quorum sensing peptides". Nucleic Acids Research. 41 (Database issue): D655–D659. doi:10.1093/nar/gks1137. PMC 3531179. PMID 23180797.
- ^ Wynendaele E, Gevaert B, Stalmans S, Verbeke F, De Spiegeleer B (September 2015). "Exploring the chemical space of quorum sensing peptides". Biopolymers. 104 (5): 544–551. doi:10.1002/bip.22649. PMID 25846138. S2CID 21031922.
- ^ O'Loughlin CT, Miller LC, Siryaporn A, Drescher K, Semmelhack MF, Bassler BL (October 2013). "A quorum-sensing inhibitor blocks Pseudomonas aeruginosa virulence and biofilm formation". Proceedings of the National Academy of Sciences of the United States of America. 110 (44): 17981–17986. Bibcode:2013PNAS..11017981O. doi:10.1073/pnas.1316981110. PMC 3816427. PMID 24143808.
- ^ Norizan SN, Yin WF, Chan KG (April 2013). "Caffeine as a potential quorum sensing inhibitor". Sensors. 13 (4): 5117–5129. Bibcode:2013Senso..13.5117N. doi:10.3390/s130405117. PMC 3673129. PMID 23598500.
- ^ a b c d Gray KM, Garey JR (August 2001). "The evolution of bacterial LuxI and LuxR quorum sensing regulators". Microbiology. 147 (Pt 8): 2379–2387. doi:10.1099/00221287-147-8-2379. PMID 11496014.
- ^ a b c Lerat E, Moran NA (May 2004). "The evolutionary history of quorum-sensing systems in bacteria". Molecular Biology and Evolution. 21 (5): 903–913. doi:10.1093/molbev/msh097. PMID 15014168.
- ^ a b Schaefer, Amy L.; Lappala, Colin R.; Morlen, Ryan P.; Pelletier, Dale A.; Lu, Tse-Yuan S.; Lankford, Patricia K.; Harwood, Caroline S.; Greenberg, E. Peter (2013-09-15). "LuxR- and LuxI-Type Quorum-Sensing Circuits Are Prevalent in Members of the Populus deltoides Microbiome". Applied and Environmental Microbiology. 79 (18): 5745–5752. Bibcode:2013ApEnM..79.5745S. doi:10.1128/AEM.01417-13. ISSN 0099-2240. PMC 3754149. PMID 23851092.
- ^ Mellbye, Brett L.; Spieck, Eva; Bottomley, Peter J.; Sayavedra-Soto, Luis A. (2017-11-15). Parales, Rebecca E. (ed.). "Acyl-Homoserine Lactone Production in Nitrifying Bacteria of the Genera Nitrosospira, Nitrobacter, and Nitrospira Identified via a Survey of Putative Quorum-Sensing Genes". Applied and Environmental Microbiology. 83 (22). Bibcode:2017ApEnM..83E1540M. doi:10.1128/AEM.01540-17. ISSN 0099-2240. PMC 5666142. PMID 28887424.
- ^ De Spiegeleer B, Verbeke F, D'Hondt M, Hendrix A, Van De Wiele C, Burvenich C, et al. (2015). "The quorum sensing peptides PhrG, CSP and EDF promote angiogenesis and invasion of breast cancer cells in vitro". PLOS ONE. 10 (3): e0119471. Bibcode:2015PLoSO..1019471D. doi:10.1371/journal.pone.0119471. PMC 4363635. PMID 25780927.
- ^ Wynendaele E, Verbeke F, D'Hondt M, Hendrix A, Van De Wiele C, Burvenich C, et al. (February 2015). "Crosstalk between the microbiome and cancer cells by quorum sensing peptides". Peptides. 64: 40–48. doi:10.1016/j.peptides.2014.12.009. hdl:2263/59248. PMID 25559405. S2CID 28064836.
- ^ Wynendaele E, Verbeke F, Stalmans S, Gevaert B, Janssens Y, Van De Wiele C, et al. (Nov 2015). "Quorum Sensing Peptides Selectively Penetrate the Blood-Brain Barrier". PLOS ONE. 10 (11): e0142071. Bibcode:2015PLoSO..1042071W. doi:10.1371/journal.pone.0142071. PMC 4633044. PMID 26536593.
- ^ a b c Bogino, P; de las Mercedes Oliva, M (July 30, 2013). "The Role of Bacterial Biofilms and Surface Components in Plant-Bacterial Associations". International Journal of Molecular Sciences. 14 (8): 15838–15859. doi:10.3390/ijms140815838. PMC 3759889. PMID 23903045.
- ^ Windsor, W. Jon (12 June 2020). "How Quorum Sensing Works". American Society for Microbiology.
- ^ Jiang, Qian; Chen, Jiashun; Yang, Chengbo; Yin, Yulong; Yao, Kang (2019-04-04). "Quorum Sensing: A Prospective Therapeutic Target for Bacterial Diseases". BioMed Research International. 2019: e2015978. doi:10.1155/2019/2015978. ISSN 2314-6133. PMC 6475571. PMID 31080810.
- ^ Zhang G, Zhang F, Ding G, Li J, Guo X, Zhu J, et al. (July 2012). "Acyl homoserine lactone-based quorum sensing in a methanogenic archaeon". The ISME Journal. 6 (7). Springer Science and Business Media LLC: 1336–1344. Bibcode:2012ISMEJ...6.1336Z. doi:10.1038/ismej.2011.203. PMC 3379639. PMID 22237544.
- ^ Callaway, Ewen (18 January 2017). "Do you speak virus? Phages caught sending chemical messages". Nature. doi:10.1038/nature.2017.21313.
- ^ Stokar-Avihail A, Tal N, Erez Z, Lopatina A, Sorek R. Widespread Utilization of Peptide Communication in Phages Infecting Soil and Pathogenic Bacteria. Cell host & microbe. 2019 May 8;25(5):746-55.
- ^ Erez Z, Steinberger-Levy I, Shamir M, Doron S, Stokar-Avihail A, Peleg Y, et al. (January 2017). "Communication between viruses guides lysis-lysogeny decisions". Nature. 541 (7638): 488–493. Bibcode:2017Natur.541..488E. doi:10.1038/nature21049. PMC 5378303. PMID 28099413.
- ^ Pierson III LS, Wood DW, Beck von Bodman S (1999). "Quorum sensing in other plant-associated bacteria". In Dunny G, Winans S (eds.). Cell-cell signaling in bacteria. Washington, D.C.: American Society for Microbiology Press. pp. 101–15. ISBN 978-1-55581-149-5. OCLC 40075829.
- ^ Watson WT, Minogue TD, Val DL, von Bodman SB, Churchill ME (March 2002). "Structural basis and specificity of acyl-homoserine lactone signal production in bacterial quorum sensing". Molecular Cell. 9 (3). Elsevier BV: 685–694. doi:10.1016/s1097-2765(02)00480-x. PMID 11931774.
- ^ Vannini A, Volpari C, Gargioli C, Muraglia E, Cortese R, De Francesco R, et al. (September 2002). "The crystal structure of the quorum sensing protein TraR bound to its autoinducer and target DNA". The EMBO Journal. 21 (17). Wiley Publishing: 4393–4401. doi:10.1093/emboj/cdf459. PMC 126196. PMID 12198141.
- ^ Zhang RG, Pappas KM, Pappas T, Brace JL, Miller PC, Oulmassov T, et al. (June 2002). "Structure of a bacterial quorum-sensing transcription factor complexed with pheromone and DNA". Nature. 417 (6892). Springer Science and Business Media LLC: 971–974. doi:10.1038/nature10294. PMID 12087407.
- ^ Götz-Rösch C, Sieper T, Fekete A, Schmitt-Kopplin P, Hartmann A, Schröder P (2015-04-10). "Influence of bacterial N-acyl-homoserine lactones on growth parameters, pigments, antioxidative capacities and the xenobiotic phase II detoxification enzymes in barley and yam bean". Frontiers in Plant Science. 6: 205. doi:10.3389/fpls.2015.00205. PMC 4392610. PMID 25914699.
- ^ Hartmann A, Schikora A (2015-08-19). "Editorial: Plant responses to bacterial quorum sensing molecules". Frontiers in Plant Science. 6: 643. doi:10.3389/fpls.2015.00643. PMC 4541025. PMID 26347761.
- ^ Von Bodman SB, Bauer WD, Coplin DL (2003-01-01). "Quorum sensing in plant-pathogenic bacteria". Annual Review of Phytopathology. 41 (1): 455–482. doi:10.1146/annurev.phyto.41.052002.095652. PMID 12730390. S2CID 3926735.
- ^ a b Alagarasan G, Aswathy KS (2017). "Shoot the Message, Not the Messenger-Combating Pathogenic Virulence in Plants by Inhibiting Quorum Sensing Mediated Signaling Molecules". Frontiers in Plant Science. 8: 556. doi:10.3389/fpls.2017.00556. PMC 5388769. PMID 28446917.
- ^ Chan KG, Atkinson S, Mathee K, Sam CK, Chhabra SR, Cámara M, et al. (January 1976). "Task delegation to physician extenders--some comparisons". American Journal of Public Health. 66 (1): 64–66. doi:10.1186/1471-2180-11-51. PMC 3062576. PMID 21385437.
- ^ Chan KG, Yin WF, Sam CK, Koh CL (February 2009). "A novel medium for the isolation of N-acylhomoserine lactone-degrading bacteria". Journal of Industrial Microbiology & Biotechnology. 36 (2): 247–251. doi:10.1007/s10295-008-0491-x. PMID 18946694. S2CID 28637950.
- ^ a b c Basavaraju M, Sisnity VS, Palaparthy R, Addanki PK (December 2016). "Quorum quenching: Signal jamming in dental plaque biofilms". Journal of Dental Sciences. 11 (4): 349–352. doi:10.1016/j.jds.2016.02.002. PMC 6395279. PMID 30894996.
- ^ Zhang LH, Dong YH (September 2004). "Quorum sensing and signal interference: diverse implications". Molecular Microbiology. 53 (6): 1563–1571. doi:10.1111/j.1365-2958.2004.04234.x. PMID 15341639. S2CID 39018931.
- ^ Manefield M, Rasmussen TB, Henzter M, Andersen JB, Steinberg P, Kjelleberg S, Givskov M (April 2002). "Halogenated furanones inhibit quorum sensing through accelerated LuxR turnover". Microbiology. 148 (Pt 4): 1119–1127. doi:10.1099/00221287-148-4-1119. PMID 11932456.
- ^ a b Chan KG, Wong CS, Yin WF, Sam CK, Koh CL (October 2010). "Rapid degradation of N-3-oxo-acylhomoserine lactones by a Bacillus cereus isolate from Malaysian rainforest soil". Antonie van Leeuwenhoek. 98 (3): 299–305. doi:10.1007/s10482-010-9438-0. PMID 20376561. S2CID 12407347.
- ^ Yoshida T, Majors RE (November 2006). "High-speed analyses using rapid resolution liquid chromatography on 1.8-microm porous particles". Journal of Separation Science. 29 (16): 2421–2432. doi:10.1002/jssc.200600267. PMID 17154122.
- ^ a b c Grandclément C, Tannières M, Moréra S, Dessaux Y, Faure D (January 2016). "Quorum quenching: role in nature and applied developments". FEMS Microbiology Reviews. 40 (1): 86–116. doi:10.1093/femsre/fuv038. PMID 26432822.
- ^ Galloway WR, Hodgkinson JT, Bowden S, Welch M, Spring DR (September 2012). "Applications of small molecule activators and inhibitors of quorum sensing in Gram-negative bacteria". Trends in Microbiology. 20 (9): 449–458. doi:10.1016/j.tim.2012.06.003. PMID 22771187.
- ^ Liu CX, Zhang DR, He Y, Zhao XS, Bai R (2010). "Modification of membrane surface for anti-biofouling performance: Effect of anti-adhesion and anti-bacteria approaches". Journal of Membrane Science. 346 (1): 121–130. doi:10.1016/j.memsci.2009.09.028.
- ^ Borea L, Naddeo V, Belgiorno V, Choo KH (December 2018). "Control of quorum sensing signals and emerging contaminants in electrochemical membrane bioreactors". Bioresource Technology. 269: 89–95. Bibcode:2018BiTec.269...89B. doi:10.1016/j.biortech.2018.08.041. PMID 30153550. S2CID 52134789.
- ^ Moradi F, Hadi N (July 2021). "Quorum-quenching activity of some Iranian medicinal plants". New Microbes and New Infections. 42: 100882. doi:10.1016/j.nmni.2021.100882. PMC 8165558. PMID 34094582.
- ^ Moradi F, Hadi N, Bazargani A (November 2020). "Evaluation of quorum-sensing inhibitory effects of extracts of three traditional medicine plants with known antibacterial properties". New Microbes and New Infections. 38: 100769. doi:10.1016/j.nmni.2020.100769. PMC 7644747. PMID 33194208.
- ^ Franks NR, Dornhaus A, Metherell BG, Nelson TR, Lanfear SA, Symes WS (January 2006). "Not everything that counts can be counted: ants use multiple metrics for a single nest trait". Proceedings. Biological Sciences. 273 (1583): 165–169. doi:10.1098/rspb.2005.3312. PMC 1560019. PMID 16555783.
- ^ Franks NR, Hooper J, Webb C, Dornhaus A (June 2005). "Tomb evaders: house-hunting hygiene in ants". Biology Letters. 1 (2): 190–192. doi:10.1098/rsbl.2005.0302. PMC 1626204. PMID 17148163.
- ^ Mallon EB, Pratt SC, et al. (2001). "Individual and collective decision-making during nest-site selection by the ant Leptothorax albipennis". Behavioral Ecology and Sociobiology. 50 (4): 352–9. Bibcode:2001BEcoS..50..352M. doi:10.1007/s002650100377. S2CID 15360262.
- ^ Pratt, S.C. (2005). "Quorum sensing by encounter rates in the ant Temnothorax albipennis". Behavioral Ecology. 16 (2): 488–96. CiteSeerX 10.1.1.598.6340. doi:10.1093/beheco/ari020.
- ^ Seeley TD, Visscher PK (2004). "Group decision making in nest-site selection by honey bees". Apidologie. 35 (2): 101–16. doi:10.1051/apido:2004004.
- ^ Seeley TD, Visscher PK (2006). "Group decision making in honey bee swarms". American Scientist. 94 (3): 220–9. doi:10.1511/2006.3.220.
- ^ You L, Cox RS, Weiss R, Arnold FH (April 2004). "Programmed population control by cell-cell communication and regulated killing". Nature. 428 (6985): 868–871. Bibcode:2004Natur.428..868Y. doi:10.1038/nature02491. PMID 15064770. S2CID 4426454.
- ^ Ma Y, Budde MW, Mayalu MN, Zhu J, Lu AC, Murray RM, Elowitz MB (March 2022). "Synthetic mammalian signaling circuits for robust cell population control". Cell. 185 (6): 967–979.e12. bioRxiv 10.1101/2020.09.02.278564. doi:10.1016/j.cell.2022.01.026. PMC 8995209. PMID 35235768. S2CID 221510088.
- ^ Hong SH, Hegde M, Kim J, Wang X, Jayaraman A, Wood TK (January 2012). "Synthetic quorum-sensing circuit to control consortial biofilm formation and dispersal in a microfluidic device". Nature Communications. 3 (1): 613. Bibcode:2012NatCo...3..613H. doi:10.1038/ncomms1616. PMC 3272573. PMID 22215088.
- ^ Din MO, Danino T, Prindle A, Skalak M, Selimkhanov J, Allen K, et al. (August 2016). "Synchronized cycles of bacterial lysis for in vivo delivery". Nature. 536 (7614): 81–85. Bibcode:2016Natur.536...81D. doi:10.1038/nature18930. PMC 5048415. PMID 27437587.
- ^ Stephens K, Pozo M, Tsao CY, Hauk P, Bentley WE (September 2019). "Bacterial co-culture with cell signaling translator and growth controller modules for autonomously regulated culture composition". Nature Communications. 10 (1): 4129. Bibcode:2019NatCo..10.4129S. doi:10.1038/s41467-019-12027-6. PMC 6739400. PMID 31511505.
- ^ a b Bassler, Bonnie L (1999-12-01). "How bacteria talk to each other: regulation of gene expression by quorum sensing". Current Opinion in Microbiology. 2 (6): 582–587. doi:10.1016/S1369-5274(99)00025-9. ISSN 1369-5274. PMID 10607620.
- ^ Britton, Matthew; Sacks, Lionel (22 August 2004). The SECOAS Project: development of a self-organising, wireless sensor network for environmental monitoring (PDF). International Workshop on Sensor and Actor Network Protocols and Applications. pp. 1–7. hdl:2440/66361. Archived from the original (PDF) on 2008-12-16.
- ^ Peysakhov M, Regli W (2005). "Ant inspired server population management in a service-based computing environment". Proceedings 2005 IEEE Swarm Intelligence Symposium, 2005. SIS 2005. Swarm Intelligence Symposium, Proceedings 2005 IEEE. pp. 357–64. doi:10.1109/SIS.2005.1501643. ISBN 0-7803-8916-6.
- ^ Sahin E, Franks N (2002). "Measurement of Space: From Ants to Robots". Proceedings of WGW 2002: EPSRC/BBSRC International Workshop. CiteSeerX 10.1.1.161.6407.
Further reading
[edit]- Dedicated issue of Philosophical Transactions B on quorum sensing (2007). Some articles are freely available.
- High citation review: Waters CM, Bassler BL (2005). "Quorum sensing: cell-to-cell communication in bacteria". Annual Review of Cell and Developmental Biology. 21: 319–346. doi:10.1146/annurev.cellbio.21.012704.131001. PMID 16212498.
External links
[edit]- The Quorum Sensing Website
- Cell-to-Cell Communication in Bacteria Archived 2013-05-17 at the Wayback Machine
- The SECOAS project—Development of a Self-Organising, Wireless Sensor Network for Environmental Monitoring
- Measurement of Space: From Ants to Robots
- Instant insight into quorum sensing from the Royal Society of Chemistry
- Bonnie Bassler: Discovering bacteria's amazing communication system Archived 2011-12-05 at the Wayback Machine
- Bonnie Bassler's seminar: "Cell-Cell Communication in Bacteria"



